RESOURCES
Putting resources into the hands of scientists, developers, and clinicians

To encourage innovation through the broad engagement of the biomedical community, CVIT shares user-friendly, well-documented, and easily accessible resources for virtual trials in medical imaging.
These resources, continuously updated, include virtual human models, the formation of digital twins, simulations of varying pathologies, simulation of imaging processes, and modeling of human and machine reading of virtual data.
Software
Graphical User Interface (GUI) for DukeSim
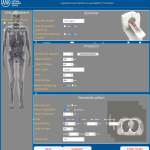 DukeSim GUI v1.0 is a webpage-based graphical user interface (GUI) to run the DukeSim CT simulator. The GUI allows users to select phantom models, scanner models, and imaging parameters to generate simulated CT images.
DukeSim GUI v1.0 is a webpage-based graphical user interface (GUI) to run the DukeSim CT simulator. The GUI allows users to select phantom models, scanner models, and imaging parameters to generate simulated CT images.TransMorph: Transformer for Unsupervised Medical Image Registration
 Novel hybrid Transformer-ConvNet model designed for 3D medical image registration.
Novel hybrid Transformer-ConvNet model designed for 3D medical image registration.CVIT Observer Models
 Toolbox for mathematical observer model calculations, designed to produce image quality figures of merit from simulated image data from CVIT.
Toolbox for mathematical observer model calculations, designed to produce image quality figures of merit from simulated image data from CVIT.Pysarfe Radiomics Pipeline
 Pipeline includes eleven unsupervised binary segmentation methods and a feature extraction module to calculate various radiomics features for an image and its binary segmentation pair.
Pipeline includes eleven unsupervised binary segmentation methods and a feature extraction module to calculate various radiomics features for an image and its binary segmentation pair.DukeSim v1.2
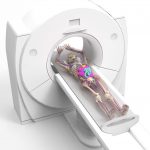 DukeSim v1.2 is a GPU-based CT simulator that generates CT projection and reconstruction images of a given voxelized computational phantom.
DukeSim v1.2 is a GPU-based CT simulator that generates CT projection and reconstruction images of a given voxelized computational phantom.XCAT Phantom Program
 Highly detailed male and female anatomies for subjects that are 50th percentile in terms of height/weight and organ volumes (thousands of defined structures including muscles and blood vessels).
Highly detailed male and female anatomies for subjects that are 50th percentile in terms of height/weight and organ volumes (thousands of defined structures including muscles and blood vessels).XCAT Brain Phantom
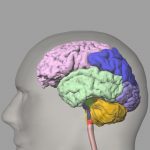 Highly detailed anatomy for the human brain, including ~100 structures and vessels. The program will generate voxelized versions of the brain at any user-defined resolution.
Highly detailed anatomy for the human brain, including ~100 structures and vessels. The program will generate voxelized versions of the brain at any user-defined resolution.Surface CT Projector
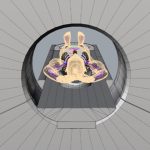 CT projector that generates x-ray projections directly from surface definitions of a given phantom without using voxelization.
CT projector that generates x-ray projections directly from surface definitions of a given phantom without using voxelization.MOBY/ROBY Phantoms
 Highly detailed anatomies for a laboratory mouse (MOBY) or laboratory rat (ROBY) with ~1400 defined structures.
Highly detailed anatomies for a laboratory mouse (MOBY) or laboratory rat (ROBY) with ~1400 defined structures.
Data
Library of Organ Dose Coefficients in Tomosynthesis Imaging
 Library of dose coefficients for organ dosimetry in tomosynthesis imaging of adults and pediatrics across diverse protocols.
Library of dose coefficients for organ dosimetry in tomosynthesis imaging of adults and pediatrics across diverse protocols.200 Anatomically Variable Adult Voxelized Phantoms
 Anatomically variable chest and abdomen phantoms, each based on CT data and modeling either COPD or lung or liver abnormalities.
Anatomically variable chest and abdomen phantoms, each based on CT data and modeling either COPD or lung or liver abnormalities.RAD-ChestCT Dataset
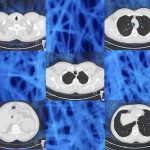 Large database of chest CT scans from unique patients. Each CT volume is annotated with a matrix of 84 abnormality labels x 52 location labels.
Large database of chest CT scans from unique patients. Each CT volume is annotated with a matrix of 84 abnormality labels x 52 location labels.2D Numerical Mouse Phantom for Dynamic Photoacoustic Tomography Studies
 Anatomically realistic spatiotemporal maps of optical absorption coefficient and photoacoustically induced pressure distribution for virtual imaging studies of dynamic photoacoustic tomography of small animal models.
Anatomically realistic spatiotemporal maps of optical absorption coefficient and photoacoustically induced pressure distribution for virtual imaging studies of dynamic photoacoustic tomography of small animal models.Multimodal Ground Truth Datasets for Abdominal Medical Image Registration
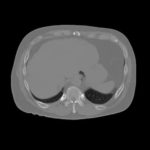 Synthesized multimodal images (T1-weighted MRI, CT, and cone beam CT) as ground truth for image segmentation and registration.
Synthesized multimodal images (T1-weighted MRI, CT, and cone beam CT) as ground truth for image segmentation and registration.Database for Benchmarking Organ Dose Estimates and Uncertainties in CT
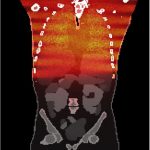 CT patient images and associated verified Monte Carlo based estimates of organ doses that may be used for benchmarking different organ dose estimation techniques against a reference standard.
CT patient images and associated verified Monte Carlo based estimates of organ doses that may be used for benchmarking different organ dose estimation techniques against a reference standard.XCAT Anatomy Files
 Library of anatomy files that work with the XCAT 2 program or can be used with other programs that work with 3D CAD geometries.
Library of anatomy files that work with the XCAT 2 program or can be used with other programs that work with 3D CAD geometries.
